- Home >
- Institut Curie News >
- A Key Protein for DNA Repair Observed in Yeast
Our DNA is naturally damaged and repaired every day. For the first time, researchers from Institute Curie led by Dr. Angela Taddei, Director of the Nucleus Dynamics Unit and Head of the Compartmentalization and Dynamics of Nuclear Functions Team, have highlighted one of the cellular mechanisms that come into play during this phenomenon. Their discovery, published in the Nature Structural & Molecular Biology journal, allows for not only observing it at work in vivo in order to understand how it works, but also in the hope of using it for future treatments targeting cancer cells.
Various environmental factors, such as ultraviolet light, or endogenous factors, such as genome replication, cause double-strand breaks in our DNA every day. However, these can lead to cell death or cause genetic instabilities that cause cancer. Fortunately, homologous recombination is one of the tools that allows our cells to repair these breaks. It consists of searching chromosomes for a sequence identical to the one that was broken in order to copy it.
"The difficulty is that the homologous sequence is embedded in thousands of sequences within the nucleus and can be several micrometers away from the one that needs to be repaired," emphasizes Dr. Angela Taddei, Director of the Nucleus Dynamics Unit (CNRS UMR3664 / Sorbonne University) and Head of the Compartmentalization and Dynamics of Nuclear Functions Team (CNRS UMR3664 / Sorbonne University). So it's not that easy to spot it... and until now, no one knew exactly how our cells worked.
A first in vivo study of Rad51
Previous studies had clearly revealed the role of a protein: Rad51. This combines with the single-stranded DNA generated at the break to form a nucleoprotein filament. But these filaments had so far only been able to be studied in vitro because the fluorescent marking of Rad51 made it non-functional. Fortunately, Dr. Angela Taddei and her team, in collaboration with Dr. Raphael Guerois of the CEA (Atomic Energy Commission) and Dr. Leonid Mirny from the Massachusetts Institute of Technology (Boston, USA), holder of a Blaise Pascal International Excellence Chair in 2021-2022, which allowed him to be welcomed at Institute Curie, have managed to overcome this obstacle.
"The key was to take into account both the structure of the protein and its evolution; the sequence of the gene coding for a specific area of Rad51 has naturally undergone numerous alterations over time, without preventing the protein from acting. So, we modified it to make the protein fluorescent, while keeping it functional," reveals Dr. Angela Taddei.
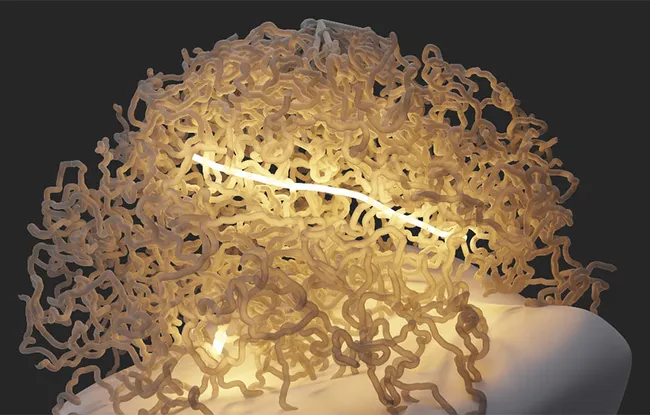
Rad51 filaments in the nucleus
This first achievement allowed the researchers to observe Rad51 in vivo with a double-strand break in baker's yeast, a unicellular organism that is simple to control. And, contrary to the common hypothesis, which suggested that the protein relies on the natural movements of the chromosomes to find the desired homologous sequence, they realized that Rad51 is very active. Because not only can the filament that it forms with single-stranded DNA reach several micrometers in length and thus be in contact simultaneously with many DNA sequences, but in addition, it also extends and contracts regularly, like a yo-yo. And each cycle of contraction and extension allows it to expand in a new direction and explore a new area of the nucleus. In two hours, the homologous sequence is identified and the repair can take place.
"Initially, we had a hard time believing what we were observing," admits Dr. Angela Taddei. "Only numerous genetic-based checks were able to convince us. The collaboration with Leonid Mirny then allowed us to demonstrate by biophysical modeling that the movements of the Rad51 filament represent an extremely effective strategy for homology search within the cell." Therefore, this discovery is really the result of the combination of structural approaches, microscopy, genetics, and biophysics.
Tumor cells "addicted" to homologous recombination
Scientists now want to understand what triggers these contraction and extension movements and how they occur, and they have already filed a patent for the application of this fundamental discovery: "Being able to observe Rad51 in vivo makes it possible for us to screen molecules to find those that disrupt its action," explains Dr. Angela Taddei. "These would be very useful against certain cancer cells, which are very fond of homologous recombination."
For this eventuality to materialize, the researchers must also demonstrate that Rad51 works in the same way in mammals and therefore create an observable version again in vivo. Which, in cells more complex than that of yeast, will inevitably be... more complex.
Reference:
S. Liu et al., In vivo Dynamics of Rad51 Filaments reveals a robust Homology Search Strategy, Nature Structural & Molecular Biology (2023).